working a sequence the place I construct mini tasks. I’ve constructed a Personal Habit and Weather Analysis mission. However I haven’t actually gotten the possibility to discover the total energy and functionality of NumPy. I wish to attempt to perceive why NumPy is so helpful in information evaluation. To wrap up this sequence, I’m going to be showcasing this in actual time.
I’ll be utilizing a fictional consumer or firm to make issues interactive. On this case, our consumer goes to be EnviroTech Dynamics, a world operator of business sensor networks.
At the moment, EnviroTech depends on outdated, loop-based Python scripts to course of over 1 million sensor readings each day. This course of is agonizingly gradual, delaying essential upkeep choices and impacting operational effectivity. They want a contemporary, high-performance resolution.
I’ve been tasked with making a NumPy-based proof-of-concept to display the best way to turbocharge their information pipeline.
The Dataset: Simulated Sensor Readings
To show the idea, I’ll be working with a big, simulated dataset generated utilizing NumPy‘s random module, that includes entries with the next key arrays:
- Temperature —Every information level represents how sizzling a machine or system element is working. These readings can shortly assist us detect when a machine begins overheating — an indication of doable failure, inefficiency, or security danger.
- Strain — information displaying how a lot strain is increase contained in the system, and whether it is inside a secure vary
- Standing codes — symbolize the well being or state of every machine or system at a given second. 0 (Regular), 1 (Warning), 2 (Vital), 3 (Defective/Lacking).
Venture Aims
The core purpose is to supply 4 clear, vectorised options to EnviroTech’s information challenges, demonstrating pace and energy. So I’ll be showcasing all of those:
- Efficiency and effectivity benchmark
- Foundational statistical baseline
- Vital anomaly detection and
- Knowledge cleansing and imputation
By the top of this text, it is best to be capable of get a full grasp of NumPy and its usefulness in information evaluation.
Goal 1: Efficiency and Effectivity Benchmark
First, we’d like a large dataset to make the pace distinction apparent. I’ll be utilizing the 1,000,000 temperature readings we deliberate earlier.
import numpy as np
# Set the dimensions of our information
NUM_READINGS = 1_000_000
# Generate the Temperature array (1 million random floating-point numbers)
# We use a seed so the outcomes are the identical each time you run the code
np.random.seed(42)
mean_temp = 45.0
std_dev_temp = 12.0
temperature_data = np.random.regular(loc=mean_temp, scale=std_dev_temp, measurement=NUM_READINGS)
print(f”Knowledge array measurement: {temperature_data.measurement} parts”)
print(f”First 5 temperatures: {temperature_data[:5]}”)Output:
Knowledge array measurement: 1000000 parts
First 5 temperatures: [50.96056984 43.34082839 52.77226246 63.27635828 42.1901595 ]Now that we’ve our data. Let’s take a look at the effectiveness of NumPy.
Assuming we needed to calculate the common of all these parts utilizing a typical Python loop, it’ll go one thing like this.
# Operate utilizing a typical Python loop
def calculate_mean_loop(information):
whole = 0
rely = 0
for worth in information:
whole += worth
rely += 1
return whole / rely
# Let’s run it as soon as to ensure it really works
loop_mean = calculate_mean_loop(temperature_data)
print(f”Imply (Loop methodology): {loop_mean:.4f}”)There’s nothing mistaken with this methodology. Nevertheless it’s fairly gradual, as a result of the pc has to course of every quantity one after the other, always shifting between the Python interpreter and the CPU.
To really showcase the pace, I’ll be utilizing the%timeit command. This runs the code a whole lot of occasions to supply a dependable common execution time.
# Time the usual Python loop (will probably be gradual)
print(“ — — Timing the Python Loop — -”)
%timeit -n 10 -r 5 calculate_mean_loop(temperature_data)Output
--- Timing the Python Loop ---
244 ms ± 51.5 ms per loop (imply ± std. dev. of 5 runs, 10 loops every)Utilizing the -n 10, I’m mainly working the code within the loop 10 occasions (to get a steady common), and utilizing the -r 5, the entire course of will probably be repeated 5 occasions (for much more stability).
Now, let’s examine this with NumPy vectorisation. And by vectorisation, it means your complete operation (common on this case) will probably be carried out on your complete array without delay, utilizing extremely optimised C code within the background.
Right here’s how the common will probably be calculated utilizing NumPy
# Utilizing the built-in NumPy imply operate
def calculate_mean_numpy(information):
return np.imply(information)
# Let’s run it as soon as to ensure it really works
numpy_mean = calculate_mean_numpy(temperature_data)
print(f”Imply (NumPy methodology): {numpy_mean:.4f}”)Output:
Imply (NumPy methodology): 44.9808Now let’s time it.
# Time the NumPy vectorized operate (will probably be quick)
print(“ — — Timing the NumPy Vectorization — -”)
%timeit -n 10 -r 5 calculate_mean_numpy(temperature_data)Output:
--- Timing the NumPy Vectorization ---
1.49 ms ± 114 μs per loop (imply ± std. dev. of 5 runs, 10 loops every)Now, that’s an enormous distinction. That’s like nearly non-existent. That’s the ability of vectorisation.
Let’s current this pace distinction to the consumer:
“We in contrast two strategies for performing the identical calculation on a million temperature readings — a conventional Python for-loop and a NumPy vectorized operation.
The distinction was dramatic: The pure Python loop took about 244 milliseconds per run whereas the NumPy model accomplished the identical process in simply 1.49 milliseconds.
That’s roughly a 160× pace enchancment.”
Goal 2: Foundational Statistical Baseline
One other cool characteristic NumPy affords is the flexibility to carry out fundamental to superior statistics — this fashion, you may get an excellent overview of what’s occurring in your dataset. It affords operations like:
- np.imply() — to calculate the common
- np.median — the center worth of the information
- np.std() — exhibits how unfold out your numbers are from the common
- np.percentile() — tells you the worth under which a sure share of your information falls.
Now that we’ve managed to supply an alternate and environment friendly resolution to retrieve and carry out summaries and calculations on their large dataset, we are able to begin taking part in round with it.
We already managed to generate our simulated temperature information. Let’s do the identical for strain. Calculating strain is a good way to display the flexibility of NumPy to deal with a number of large arrays very quickly in any respect.
For our consumer, it additionally permits me to showcase a well being test on their industrial programs.
Additionally, temperature and strain are sometimes associated. A sudden strain drop may be the reason for a spike in temperature, or vice versa. Calculating baselines for each permits us to see if they’re drifting collectively or independently
# Generate the Strain array (Uniform distribution between 100.0 and 500.0)
np.random.seed(43) # Use a unique seed for a brand new dataset
pressure_data = np.random.uniform(low=100.0, excessive=500.0, measurement=1_000_000)
print(“Knowledge arrays prepared.”)Output:
Knowledge arrays prepared.Alright, let’s start our calculations.
print(“n — — Temperature Statistics — -”)
# 1. Imply and Median
temp_mean = np.imply(temperature_data)
temp_median = np.median(temperature_data)
# 2. Customary Deviation
temp_std = np.std(temperature_data)
# 3. Percentiles (Defining the 90% Regular Vary)
temp_p5 = np.percentile(temperature_data, 5) # fifth percentile
temp_p95 = np.percentile(temperature_data, 95) # ninety fifth percentile
# Formating our outcomes
print(f”Imply (Common): {temp_mean:.2f}°C”)
print(f”Median (Center): {temp_median:.2f}°C”)
print(f”Std. Deviation (Unfold): {temp_std:.2f}°C”)
print(f”90% Regular Vary: {temp_p5:.2f}°C to {temp_p95:.2f}°C”)Right here’s the output:
--- Temperature Statistics ---
Imply (Common): 44.98°C
Median (Center): 44.99°C
Std. Deviation (Unfold): 12.00°C
90% Regular Vary: 25.24°C to 64.71°CSo to elucidate what you’re seeing right here
The Imply (Common): 44.98°C mainly provides us a central level round which most readings are anticipated to fall. That is fairly cool as a result of we don’t must scan via your complete massive dataset. With this quantity, I’ve gotten a fairly good thought of the place our temperature readings normally fall.
The Median (Center): 44.99°C is sort of an identical to the imply if you happen to discover. This tells us that there aren’t excessive outliers dragging the common too excessive or too low.
The usual deviation of 12°C means the temperatures fluctuate fairly a bit from the common. Mainly, some days are a lot hotter or cooler than others. A decrease worth (say 3°C or 4°C) would have prompt extra consistency, however 12°C signifies a extremely variable sample.
For the percentile, it mainly means most days hover between 25°C and 65°C,
If I had been to current this to the consumer, I may put it like this:
“On common, the system (or setting) maintains a temperature round 45°C, which serves as a dependable baseline for typical working or environmental circumstances. A deviation of 12°C signifies that temperature ranges fluctuate considerably across the common.
To place it merely, the readings aren’t very steady. Lastly, 90% of all readings fall between 25°C and 65°C. This offers a sensible image of what “regular” seems like, serving to you outline acceptable thresholds for alerts or upkeep. To enhance efficiency or reliability, we may establish the causes of excessive fluctuations (e.g., exterior warmth sources, air flow patterns, system load).”
Let’s calculate for strain additionally.
print(“n — — Strain Statistics — -”)
# Calculate all 5 measures for Strain
pressure_stats = {
“Imply”: np.imply(pressure_data),
“Median”: np.median(pressure_data),
“Std. Dev”: np.std(pressure_data),
“fifth %tile”: np.percentile(pressure_data, 5),
“ninety fifth %tile”: np.percentile(pressure_data, 95),
}
for label, worth in pressure_stats.gadgets():
print(f”{label:<12}: {worth:.2f} kPa”)To enhance our codebase, I’m storing all of the calculations carried out in a dictionary known as strain stats, and I’m merely looping over the key-value pairs.
Right here’s the output:
--- Strain Statistics ---
Imply : 300.09 kPa
Median : 300.04 kPa
Std. Dev : 115.47 kPa
fifth %tile : 120.11 kPa
ninety fifth %tile : 480.09 kPaIf I had been to current this to the consumer. It’d go one thing like this:
“Our strain readings common round 300 kilopascals, and the median — the center worth — is sort of the identical. That tells us the strain distribution is sort of balanced general. Nonetheless, the customary deviation is about 115 kPa, which suggests there’s loads of variation between readings. In different phrases, some readings are a lot greater or decrease than the everyday 300 kPa stage.
Wanting on the percentiles, 90% of our readings fall between 120 and 480 kPa. That’s a variety, suggesting that strain circumstances aren’t steady — presumably fluctuating between high and low states throughout operation. So whereas the common seems fantastic, the variability may level to inconsistent efficiency or environmental components affecting the system.”
Goal 3: Vital Anomaly Identification
Considered one of my favorite options of NumPy is the flexibility to shortly establish and filter out anomalies in your dataset. To display this, our fictional consumer, EnviroTech Dynamics, offered us with one other useful array that incorporates system standing codes. This tells us how the machine is constantly working. It’s merely a variety of codes (0–3).
- 0 → Regular
- 1 → Warning
- 2 → Vital
- 3 → Sensor Error
They obtain hundreds of thousands of readings per day, and our job is to seek out each machine that’s each in a essential state and working dangerously sizzling.
Doing this manually, and even with a loop, would take ages. That is the place Boolean Indexing (masking) is available in. It lets us filter large datasets in milliseconds by making use of logical circumstances on to arrays, with out loops.
Earlier, we generated our temperature and strain information. Let’s do the identical for the standing codes.
# Reusing 'temperature_data' from earlier
import numpy as np
np.random.seed(42) # For reproducibility
status_codes = np.random.selection(
a=[0, 1, 2, 3],
measurement=len(temperature_data),
p=[0.85, 0.10, 0.03, 0.02] # 0=Regular, 1=Warning, 2=Vital, 3=Offline
)
# Let’s preview our information
print(status_codes[:5])Output:
[0 2 0 0 0]Every temperature studying now has an identical standing code. This permits us to pinpoint which sensors report issues and how extreme they’re.
Subsequent, we’ll want some form of threshold or anomaly standards. In most situations, something above imply + 3 × customary deviation is taken into account a extreme outlier, the sort of studying you don’t need in your system. To compute that
temp_mean = np.imply(temperature_data)
temp_std = np.std(temperature_data)
SEVERITY_THRESHOLD = temp_mean + (3 * temp_std)
print(f”Extreme Outlier Threshold: {SEVERITY_THRESHOLD:.2f}°C”)Output:
Extreme Outlier Threshold: 80.99°CSubsequent, we’ll create two filters (masks) to isolate information that meets our circumstances. One for readings the place the system standing is Vital (code 2) and one other for readings the place the temperature exceeds the brink.
# Masks 1 — Readings the place system standing = Vital (code 2)
critical_status_mask = (status_codes == 2)
# Masks 2 — Readings the place temperature exceeds threshold
high_temp_outlier_mask = (temperature_data > SEVERITY_THRESHOLD)
print(f”Vital standing readings: {critical_status_mask.sum()}”)
print(f”Excessive-temp outliers: {high_temp_outlier_mask.sum()}”)Right here’s what’s occurring behind the scenes. NumPy creates two arrays crammed with True or False. Each True marks a studying that satisfies the situation. True will probably be represented as 1, and False will probably be represented as 0. Summing them shortly counts what number of match.
Right here’s the output:
Vital standing readings: 30178
Excessive-temp outliers: 1333Let’s mix each anomalies earlier than printing our remaining end result. We would like readings which are each essential and too sizzling. NumPy permits us to filter on a number of circumstances utilizing logical operators. On this case, we’ll be utilizing the AND operate represented as &.
# Mix each circumstances with a logical AND
critical_anomaly_mask = critical_status_mask & high_temp_outlier_mask
# Extract precise temperatures of these anomalies
extracted_anomalies = temperature_data[critical_anomaly_mask]
anomaly_count = critical_anomaly_mask.sum()
print(“n — — Last Outcomes — -”)
print(f”Complete Vital Anomalies: {anomaly_count}”)
print(f”Pattern Temperatures: {extracted_anomalies[:5]}”)Output:
--- Last Outcomes ---
Complete Vital Anomalies: 34
Pattern Temperatures: [81.9465697 81.11047892 82.23841531 86.65859372 81.146086 ]Let’s current this to the consumer
“After analyzing a million temperature readings, our system detected 34 essential anomalies — readings that had been each flagged as ‘essential standing’ by the machine and exceeded the high-temperature threshold.
The primary few of those readings fall between 81°C and 86°C, which is properly above our regular working vary of round 45°C. This implies {that a} small variety of sensors are reporting harmful spikes, presumably indicating overheating or sensor malfunction.
In different phrases, whereas 99.99% of our information seems steady, these 34 factors symbolize the precise spots the place we should always focus upkeep or examine additional.”
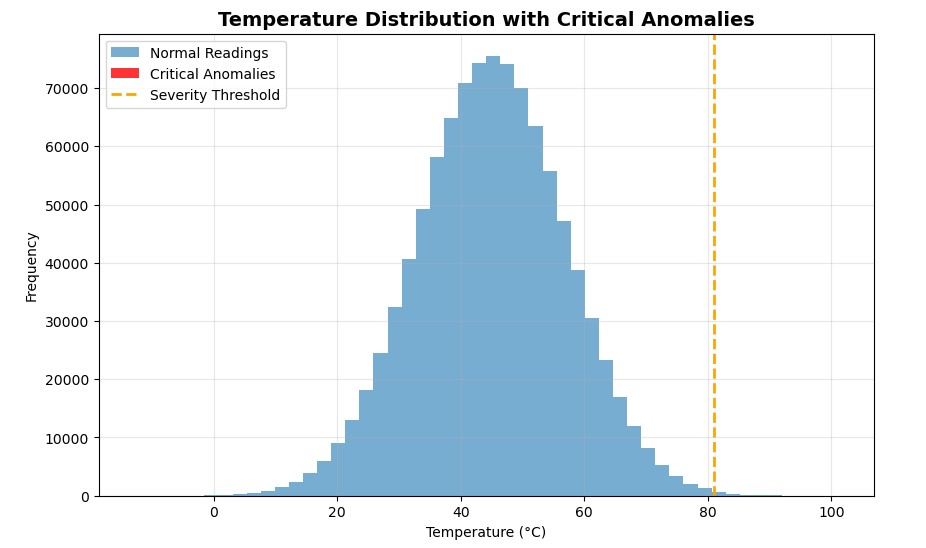
Let’s visualise this actual fast with matplotlib
After I first plotted the outcomes, I anticipated to see a cluster of purple bars displaying my essential anomalies. However there have been none.
At first, I assumed one thing was mistaken, however then it clicked. Out of 1 million readings, solely 34 had been essential. That’s the fantastic thing about Boolean masking: it detects what your eyes can’t. Even when the anomalies disguise deep inside hundreds of thousands of regular values, NumPy flags them in milliseconds.
Goal 4: Knowledge Cleansing and Imputation
Lastly, NumPy lets you do away with inconsistencies and information that doesn’t make sense. You might need come throughout the idea of information cleansing in information evaluation. In Python, NumPy and Pandas are sometimes used to streamline this exercise.
To display this, our status_codes comprise entries with a price of three (Defective/Lacking). If we use these defective temperature readings in our general evaluation, they are going to skew our outcomes. The answer is to interchange the defective readings with a statistically sound estimated worth.
Step one is to determine what worth we should always use to interchange the dangerous information. The median is all the time an ideal selection as a result of, in contrast to the imply, it’s much less affected by excessive values.
# TASK: Determine the masks for ‘Legitimate’ information (the place status_codes is NOT 3 — Defective/Lacking).
valid_data_mask = (status_codes != 3)
# TASK: Calculate the median temperature ONLY for the Legitimate information factors. That is our imputation worth.
valid_median_temp = np.median(temperature_data[valid_data_mask])
print(f”Median of all legitimate readings: {valid_median_temp:.2f}°C”)Output:
Median of all legitimate readings: 44.99°CNow, we’ll carry out some conditional substitute utilizing the highly effective np.the place() operate. Right here’s a typical construction of the operate.
np.the place(Situation, Value_if_True, Value_if_False)
In our case:
- Situation: Is the standing code 3 (Defective/Lacking)?
- Worth if True: Use our calculated
valid_median_temp. - Worth if False: Maintain the unique temperature studying.
# TASK: Implement the conditional substitute utilizing np.the place().
cleaned_temperature_data = np.the place(
status_codes == 3, # CONDITION: Is the studying defective?
valid_median_temp, # VALUE_IF_TRUE: Substitute with the calculated median.
temperature_data # VALUE_IF_FALSE: Maintain the unique temperature worth.
)
# TASK: Print the entire variety of changed values.
imputed_count = (status_codes == 3).sum()
print(f”Complete Defective readings imputed: {imputed_count}”)Output:
Complete Defective readings imputed: 20102I didn’t count on the lacking values to be this a lot. It in all probability affected our studying above ultimately. Good factor, we managed to interchange them in seconds.
Now, let’s confirm the repair by checking the median for each the unique and cleaned information
# TASK: Print the change within the general imply or median to point out the impression of the cleansing.
print(f”nOriginal Median: {np.median(temperature_data):.2f}°C”)
print(f”Cleaned Median: {np.median(cleaned_temperature_data):.2f}°C”)Output:
Authentic Median: 44.99°C
Cleaned Median: 44.99°COn this case, even after cleansing over 20,000 defective data, the median temperature remained regular at 44.99°C, indicating that the dataset is statistically sound and balanced.
Let’s current this to the consumer:
“Out of 1 million temperature readings, 20,102 had been marked as defective (standing code = 3). As a substitute of eradicating these defective data, we changed them with the median temperature worth (≈ 45°C) — a typical data-cleaning method that retains the dataset constant with out distorting the development.
Curiously, the median temperature remained unchanged (44.99°C) earlier than and after cleansing. That’s an excellent signal: it means the defective readings didn’t skew the dataset, and the substitute didn’t alter the general information distribution.”
Conclusion
And there we go! We initiated this mission to deal with a essential difficulty for EnviroTech Dynamics: the necessity for sooner, loop-free information evaluation. The facility of NumPy arrays and vectorisation allowed us to repair the issue and future-proof their analytical pipeline.
NumPy ndarray is the silent engine of your complete Python information science ecosystem. Each main library, like Pandas, scikit-learn, TensorFlow, and PyTorch, makes use of NumPy arrays at its core for quick numerical computation.
By mastering NumPy, you’ve constructed a robust analytical basis. The subsequent logical step for me is to maneuver from single arrays to structured evaluation with the Pandas library, which organises NumPy arrays into tables (DataFrames) for even simpler labelling and manipulation.
Thanks for studying! Be at liberty to attach with me: